This tutorial depends on step-16.
This program was contributed by Thomas C. Clevenger and Timo Heister.
The creation of this tutorial was partially supported by NSF Award DMS-1522191, DMS-1901529, OAC-1835452, by the Computational Infrastructure in Geodynamics initiative (CIG), through the NSF under Award EAR-0949446 and EAR-1550901 and The University of California - Davis.
- Note
- If you use this program as a basis for your own work, please consider citing it in your list of references. The initial version of this work was contributed to the deal.II project by the authors listed in the following citation:

Introduction
This program solves an advection-diffusion problem using a geometric multigrid (GMG) preconditioner. The basics of this preconditioner are discussed in step-16; here we discuss the necessary changes needed for a non-symmetric PDE. Additionally, we introduce the idea of block smoothing (as compared to point smoothing in step-16), and examine the effects of DoF renumbering for additive and multiplicative smoothers.
Equation
The advection-diffusion equation is given by
\begin{align*}
-\varepsilon \Delta u + \boldsymbol{\beta}\cdot \nabla u & = f &
\text{ in } \Omega\\
u &= g & \text{ on } \partial\Omega
\end{align*}
where \varepsilon>0, \boldsymbol{\beta} is the advection direction, and f is a source. A few notes:
- If \boldsymbol{\beta}=\boldsymbol{0}, this is the Laplace equation solved in step-16 (and many other places).
- If \varepsilon=0 then this is the stationary advection equation solved in step-9.
- One can define a dimensionless number for this problem, called the Peclet number: \mathcal{P} \dealcoloneq \frac{\|\boldsymbol{\beta}\|
L}{\varepsilon}, where L is the length scale of the domain. It characterizes the kind of equation we are considering: If \mathcal{P}>1, we say the problem is advection-dominated, else if \mathcal{P}<1 we will say the problem is diffusion-dominated.
For the discussion in this tutorial we will be concerned with advection-dominated flow. This is the complicated case: We know that for diffusion-dominated problems, the standard Galerkin method works just fine, and we also know that simple multigrid methods such as those defined in step-16 are very efficient. On the other hand, for advection-dominated problems, the standard Galerkin approach leads to oscillatory and unstable discretizations, and simple solvers are often not very efficient. This tutorial program is therefore intended to address both of these issues.
Streamline diffusion
Using the standard Galerkin finite element method, for suitable test functions v_h, a discrete weak form of the PDE would read
\begin{align*}
a(u_h,v_h) = F(v_h)
\end{align*}
where
\begin{align*}
a(u_h,v_h) &= (\varepsilon \nabla v_h,\, \nabla u_h) +
(v_h,\,\boldsymbol{\beta}\cdot \nabla u_h),\\
F(v_h) &= (v_h,\,f).
\end{align*}
Unfortunately, one typically gets oscillatory solutions with this approach. Indeed, the following error estimate can be shown for this formulation:
\begin{align*}
\|\nabla (u-u_h)\| \leq (1+\mathcal{P}) \inf_{v_h} \|\nabla (u-v_h)\|.
\end{align*}
The infimum on the right can be estimated as follows if the exact solution is sufficiently smooth:
\begin{align*}
\inf_{v_h} \|\nabla (u-v_h)\|.
\le
\|\nabla (u-I_h u)\|
\le
h^k
C
\|\nabla^k u)\|
\end{align*}
where k is the polynomial degree of the finite elements used. As a consequence, we obtain the estimate
\begin{align*}
\|\nabla (u-u_h)\|
\leq (1+\mathcal{P}) C h^k
\|\nabla^k u)\|.
\end{align*}
In other words, the numerical solution will converge. On the other hand, given the definition of \mathcal{P} above, we have to expect poor numerical solutions with a large error when \varepsilon \ll
\|\boldsymbol{\beta}\| L, i.e., if the problem has only a small amount of diffusion.
To combat this, we will consider the new weak form
\begin{align*}
a(u_h,\,v_h) + \sum_K (-\varepsilon \Delta u_h +
\boldsymbol{\beta}\cdot \nabla u_h-f,\,\delta_K
\boldsymbol{\beta}\cdot \nabla v_h)_K = F(v_h)
\end{align*}
where the sum is done over all cells K with the inner product taken for each cell, and \delta_K is a cell-wise constant stabilization parameter defined in [69].
Essentially, adding in the discrete strong form residual enhances the coercivity of the bilinear form a(\cdot,\cdot) which increases the stability of the discrete solution. This method is commonly referred to as streamline diffusion or SUPG (streamline upwind/Petrov-Galerkin).
Smoothers
One of the goals of this tutorial is to expand from using a simple (point-wise) Gauss-Seidel (SOR) smoother that is used in step-16 (class PreconditionSOR) on each level of the multigrid hierarchy. The term "point-wise" is traditionally used in solvers to indicate that one solves at one "grid point" at a time; for scalar problems, this means to use a solver that updates one unknown of the linear system at a time, keeping all of the others fixed; one would then iterate over all unknowns in the problem and, once done, start over again from the first unknown until these "sweeps" converge. Jacobi, Gauss-Seidel, and SOR iterations can all be interpreted in this way. In the context of multigrid, one does not think of these methods as "solvers", but as "smoothers". As such, one is not interested in actually solving the linear system. It is enough to remove the high-frequency part of the residual for the multigrid method to work, because that allows restricting the solution to a coarser mesh. Therefore, one only does a few, fixed number of "sweeps" over all unknowns. In the code in this tutorial this is controlled by the "Smoothing steps" parameter.
But these methods are known to converge rather slowly when used as solvers. While as multigrid smoothers, they are surprisingly good, they can also be improved upon. In particular, we consider "cell-based" smoothers here as well. These methods solve for all unknowns on a cell at once, keeping all other unknowns fixed; they then move on to the next cell, and so on and so forth. One can think of them as "block" versions of Jacobi, Gauss-Seidel, or SOR, but because degrees of freedom are shared among multiple cells, these blocks overlap and the methods are in fact best be explained within the framework of additive and multiplicative Schwarz methods.
In contrast to step-16, our test problem contains an advective term. Especially with a small diffusion constant \varepsilon, information is transported along streamlines in the given advection direction. This means that smoothers are likely to be more effective if they allow information to travel in downstream direction within a single smoother application. If we want to solve one unknown (or block of unknowns) at a time in the order in which these unknowns (or blocks) are enumerated, then this information propagation property requires reordering degrees of freedom or cells (for the cell-based smoothers) accordingly so that the ones further upstream are treated earlier (have lower indices) and those further downstream are treated later (have larger indices). The influence of the ordering will be visible in the results section.
Let us now briefly define the smoothers used in this tutorial. For a more detailed introduction, we refer to [72] and the books [110] and [114]. A Schwarz preconditioner requires a decomposition
\begin{align*}
V = \sum_{j=1}^J V_j
\end{align*}
of our finite element space V. Each subproblem V_j also has a Ritz projection P_j: V \rightarrow V_j based on the bilinear form a(\cdot,\cdot). This projection induces a local operator A_j for each subproblem V_j. If \Pi_j:V\rightarrow V_j is the orthogonal projector onto V_j, one can show A_jP_j=\Pi_j^TA.
With this we can define an additive Schwarz preconditioner for the operator A as
\begin{align*}
B^{-1} = \sum_{j=1}^J P_j A^{-1} = \sum_{j=1}^J A_j^{-1} \Pi_j^T.
\end{align*}
In other words, we project our solution into each subproblem, apply the inverse of the subproblem A_j, and sum the contributions up over all j.
Note that one can interpret the point-wise (one unknown at a time) Jacobi method as an additive Schwarz method by defining a subproblem V_j for each degree of freedom. Then, A_j^{-1} becomes a multiplication with the inverse of a diagonal entry of A.
For the "Block Jacobi" method used in this tutorial, we define a subproblem V_j for each cell of the mesh on the current level. Note that we use a continuous finite element, so these blocks are overlapping, as degrees of freedom on an interface between two cells belong to both subproblems. The logic for the Schwarz operator operating on the subproblems (in deal.II they are called "blocks") is implemented in the class RelaxationBlock. The "Block
Jacobi" method is implemented in the class RelaxationBlockJacobi. Many aspects of the class (for example how the blocks are defined and how to invert the local subproblems A_j) can be configured in the smoother data, see RelaxationBlock::AdditionalData and DoFTools::make_cell_patches() for details.
So far, we discussed additive smoothers where the updates can be applied independently and there is no information flowing within a single smoother application. A multiplicative Schwarz preconditioner addresses this and is defined by
\begin{align*}
B^{-1} = \left( I- \prod_{j=1}^J \left(I-P_j\right) \right) A^{-1}.
\end{align*}
In contrast to above, the updates on the subproblems V_j are applied sequentially. This means that the update obtained when inverting the subproblem A_j is immediately used in A_{j+1}. This becomes visible when writing out the project:
\begin{align*}
B^{-1}
=
\left(
I
-
\left(I-P_1\right)\left(I-P_2\right)\cdots\left(I-P_J\right)
\right)
A^{-1}
=
A^{-1}
-
\left[ \left(I-P_1\right)
\left[ \left(I-P_2\right)\cdots
\left[\left(I-P_J\right) A^{-1}\right] \cdots \right] \right]
\end{align*}
When defining the sub-spaces V_j as whole blocks of degrees of freedom, this method is implemented in the class RelaxationBlockSOR and used when you select "Block SOR" in this tutorial. The class RelaxationBlockSOR is also derived from RelaxationBlock. As such, both additive and multiplicative Schwarz methods are implemented in a unified framework.
Finally, let us note that the standard Gauss-Seidel (or SOR) method can be seen as a multiplicative Schwarz method with a subproblem for each DoF.
Test problem
We will be considering the following test problem: \Omega =
[-1,\,1]\times[-1,\,1]\backslash B_{0.3}(0), i.e., a square with a circle of radius 0.3 centered at the origin removed. In addition, we use \varepsilon=0.005, \boldsymbol{\beta} =
[-\sin(\pi/6),\,\cos(\pi/6)], f=0, and Dirichlet boundary values
\begin{align*}
g = \left\{\begin{array}{ll} 1 & \text{if } x=-1 \text{ or } y=-1,\,x\geq 0.5 \\
0 & \text{otherwise} \end{array}\right.
\end{align*}
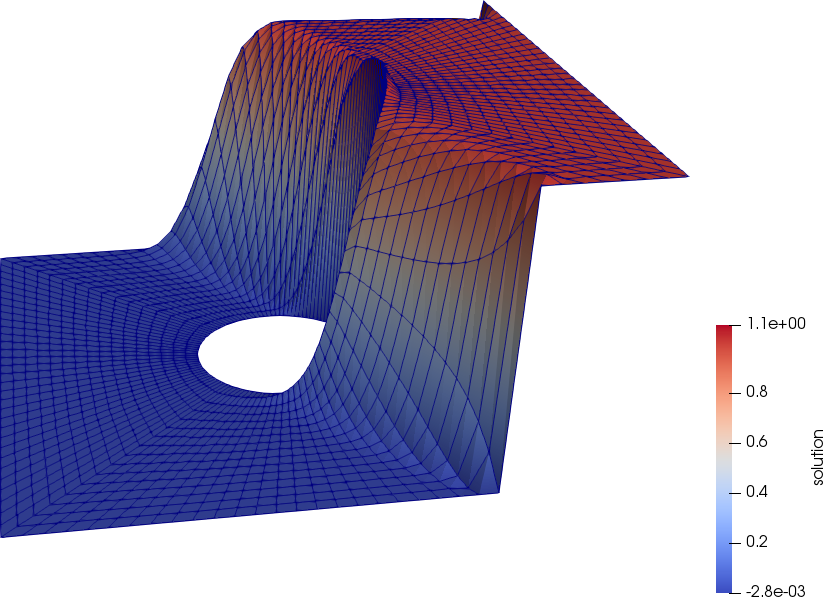
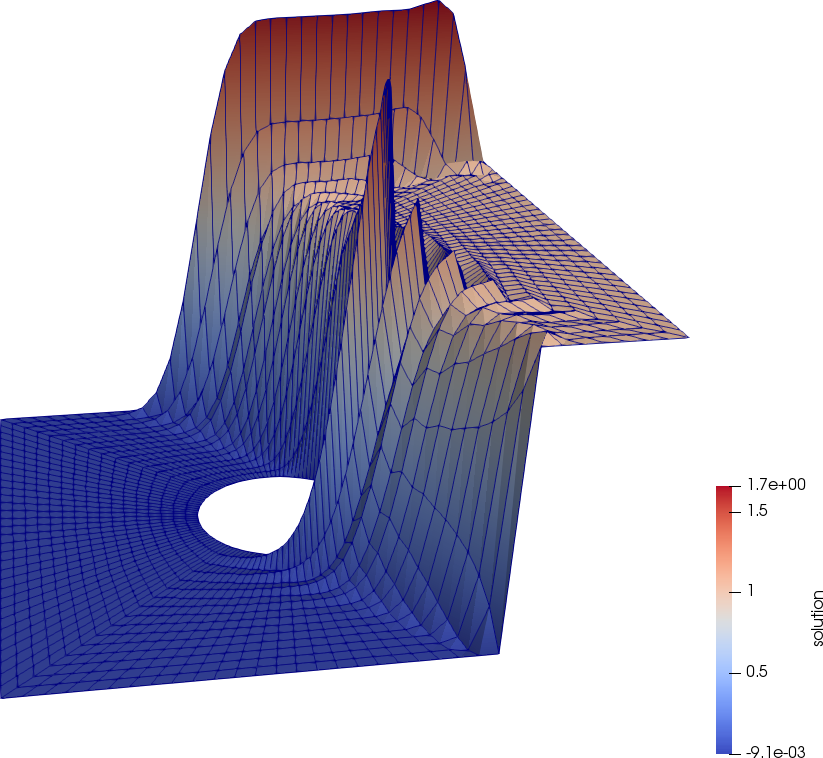
The following figures depict the solutions with (left) and without (right) streamline diffusion. Without streamline diffusion we see large oscillations around the boundary layer, demonstrating the instability of the standard Galerkin finite element method for this problem.
The commented program
Include files
Typical files needed for standard deal.II:
Include all relevant multilevel files:
C++:
#include <algorithm>
#include <fstream>
#include <iostream>
#include <random>
We will be using MeshWorker::mesh_loop functionality for assembling matrices:
As always, we will be putting everything related to this program into a namespace of its own.
Since we will be using the MeshWorker framework, the first step is to define the following structures needed by the assemble_cell() function used by MeshWorker::mesh_loop(): ScratchData contains an FEValues object which is needed for assembling a cell's local contribution, while CopyData contains the output from a cell's local contribution and necessary information to copy that to the global system. (Their purpose is also explained in the documentation of the WorkStream class.)
namespace Step63
{
template <int dim>
struct ScratchData
{
const unsigned int quadrature_degree)
: fe_values(fe,
QGauss<dim>(quadrature_degree),
{}
ScratchData(const ScratchData<dim> &scratch_data)
: fe_values(scratch_data.fe_values.get_fe(),
scratch_data.fe_values.get_quadrature(),
{}
};
struct CopyData
{
CopyData() = default;
unsigned int dofs_per_cell;
std::vector<types::global_dof_index> local_dof_indices;
};
@ update_hessians
Second derivatives of shape functions.
@ update_values
Shape function values.
@ update_JxW_values
Transformed quadrature weights.
@ update_gradients
Shape function gradients.
@ update_quadrature_points
Transformed quadrature points.
void cell_matrix(FullMatrix< double > &M, const FEValuesBase< dim > &fe, const FEValuesBase< dim > &fetest, const ArrayView< const std::vector< double > > &velocity, const double factor=1.)
Problem parameters
The second step is to define the classes that deal with run-time parameters to be read from an input file.
We will use ParameterHandler to pass in parameters at runtime. The structure Settings parses and stores the parameters to be queried throughout the program.
{
enum DoFRenumberingStrategy
{
upstream,
};
void get_parameters(const std::string &prm_filename);
unsigned int fe_degree;
std::string smoother_type;
unsigned int smoothing_steps;
DoFRenumberingStrategy dof_renumbering;
bool with_streamline_diffusion;
bool output;
};
void Settings::get_parameters(const std::string &prm_filename)
{
"0.005",
"Diffusion parameter");
"1",
"Finite Element degree");
"block SOR",
"Select smoother: SOR|Jacobi|block SOR|block Jacobi");
"2",
"Number of smoothing steps");
"DoF renumbering",
"downstream",
"Select DoF renumbering: none|downstream|upstream|random");
"true",
"Enable streamline diffusion stabilization: true|false");
"true",
"Generate graphical output: true|false");
if (prm_filename.empty())
{
false,
ExcMessage(
"Please pass a .prm file as the first argument!"));
}
smoother_type = prm.
get(
"Smoother type");
const std::string renumbering = prm.
get(
"DoF renumbering");
if (renumbering == "none")
else if (renumbering == "downstream")
else if (renumbering == "upstream")
dof_renumbering = DoFRenumberingStrategy::upstream;
else if (renumbering == "random")
else
"an invalid value."));
with_streamline_diffusion = prm.
get_bool(
"With streamline diffusion");
}
virtual void parse_input(std::istream &input, const std::string &filename="input file", const std::string &last_line="", const bool skip_undefined=false)
std::ostream & print_parameters(std::ostream &out, const OutputStyle style) const
long int get_integer(const std::string &entry_string) const
bool get_bool(const std::string &entry_name) const
void declare_entry(const std::string &entry, const std::string &default_value, const Patterns::PatternBase &pattern=Patterns::Anything(), const std::string &documentation="", const bool has_to_be_set=false)
std::string get(const std::string &entry_string) const
double get_double(const std::string &entry_name) const
static ::ExceptionBase & ExcMessage(std::string arg1)
#define AssertThrow(cond, exc)
void downstream(DoFHandler< dim, spacedim > &dof_handler, const Tensor< 1, spacedim > &direction, const bool dof_wise_renumbering=false)
void random(DoFHandler< dim, spacedim > &dof_handler)
SymmetricTensor< 2, dim, Number > epsilon(const Tensor< 2, dim, Number > &Grad_u)
Cell permutations
The ordering in which cells and degrees of freedom are traversed will play a role in the speed of convergence for multiplicative methods. Here we define functions which return a specific ordering of cells to be used by the block smoothers.
For each type of cell ordering, we define a function for the active mesh and one for a level mesh (i.e., for the cells at one level of a multigrid hierarchy). While the only reordering necessary for solving the system will be on the level meshes, we include the active reordering for visualization purposes in output_results().
For the two downstream ordering functions, we first create an array with all of the relevant cells that we then sort in downstream direction using a "comparator" object. The output of the functions is then simply an array of the indices of the cells in the just computed order.
template <int dim>
std::vector<unsigned int>
const unsigned int level)
{
std::vector<typename DoFHandler<dim>::level_cell_iterator> ordered_cells;
ordered_cells.push_back(cell);
const DoFRenumbering::
CompareDownstream<typename DoFHandler<dim>::level_cell_iterator, dim>
comparator(direction);
std::sort(ordered_cells.begin(), ordered_cells.end(), comparator);
std::vector<unsigned> ordered_indices;
for (const auto &cell : ordered_cells)
ordered_indices.push_back(cell->index());
return ordered_indices;
}
template <int dim>
std::vector<unsigned int>
{
std::vector<typename DoFHandler<dim>::active_cell_iterator> ordered_cells;
ordered_cells.push_back(cell);
const DoFRenumbering::
CompareDownstream<typename DoFHandler<dim>::active_cell_iterator, dim>
comparator(direction);
std::sort(ordered_cells.begin(), ordered_cells.end(), comparator);
std::vector<unsigned int> ordered_indices;
for (const auto &cell : ordered_cells)
ordered_indices.push_back(cell->index());
return ordered_indices;
}
const Triangulation< dim, spacedim > & get_triangulation() const
unsigned int n_active_cells() const
unsigned int n_cells() const
IteratorRange< active_cell_iterator > active_cell_iterators() const
IteratorRange< cell_iterator > cell_iterators_on_level(const unsigned int level) const
The functions that produce a random ordering are similar in spirit in that they first put information about all cells into an array. But then, instead of sorting them, they shuffle the elements randomly using the facilities C++ offers to generate random numbers. The way this is done is by iterating over all elements of the array, drawing a random number for another element before that, and then exchanging these elements. The result is a random shuffle of the elements of the array.
template <int dim>
std::vector<unsigned int>
const unsigned int level)
{
std::vector<unsigned int> ordered_cells;
ordered_cells.push_back(cell->index());
std::mt19937 random_number_generator;
std::shuffle(ordered_cells.begin(),
ordered_cells.end(),
random_number_generator);
return ordered_cells;
}
template <int dim>
std::vector<unsigned int>
{
std::vector<unsigned int> ordered_cells;
ordered_cells.push_back(cell->index());
std::mt19937 random_number_generator;
std::shuffle(ordered_cells.begin(),
ordered_cells.end(),
random_number_generator);
return ordered_cells;
}
Right-hand side and boundary values
The problem solved in this tutorial is an adaptation of Ex. 3.1.3 found on pg. 118 of Finite Elements and Fast Iterative Solvers: with Applications in Incompressible Fluid Dynamics by Elman, Silvester, and Wathen. The main difference being that we add a hole in the center of our domain with zero Dirichlet boundary conditions.
For a complete description, we need classes that implement the zero right-hand side first (we could of course have just used Functions::ZeroFunction):
template <int dim>
class RightHandSide :
public Function<dim>
{
public:
const unsigned int component = 0) const override;
const unsigned int component = 0) const override;
};
template <int dim>
double RightHandSide<dim>::value(
const Point<dim> &,
const unsigned int component) const
{
(void)component;
return 0.0;
}
template <int dim>
void RightHandSide<dim>::value_list(
const std::vector<
Point<dim>> &points,
const unsigned int component) const
{
for (unsigned int i = 0; i < points.size(); ++i)
values[i] = RightHandSide<dim>::value(points[i], component);
}
virtual void value_list(const std::vector< Point< dim > > &points, std::vector< RangeNumberType > &values, const unsigned int component=0) const
virtual RangeNumberType value(const Point< dim > &p, const unsigned int component=0) const
static ::ExceptionBase & ExcIndexRange(int arg1, int arg2, int arg3)
#define Assert(cond, exc)
static ::ExceptionBase & ExcDimensionMismatch(std::size_t arg1, std::size_t arg2)
We also have Dirichlet boundary conditions. On a connected portion of the outer, square boundary we set the value to 1, and we set the value to 0 everywhere else (including the inner, circular boundary):
template <int dim>
class BoundaryValues :
public Function<dim>
{
public:
const unsigned int component = 0) const override;
const unsigned int component = 0) const override;
};
template <int dim>
double BoundaryValues<dim>::value(
const Point<dim> & p,
const unsigned int component) const
{
(void)component;
Set boundary to 1 if x=1, or if x>0.5 and y=-1.
{
return 1.0;
}
else
{
return 0.0;
}
}
template <int dim>
void BoundaryValues<dim>::value_list(
const std::vector<
Point<dim>> &points,
const unsigned int component) const
{
for (unsigned int i = 0; i < points.size(); ++i)
values[i] = BoundaryValues<dim>::value(points[i], component);
}
Expression fabs(const Expression &x)
SymmetricTensor< 2, dim, Number > e(const Tensor< 2, dim, Number > &F)
Streamline diffusion implementation
The streamline diffusion method has a stabilization constant that we need to be able to compute. The choice of how this parameter is computed is taken from On Discontinuity-Capturing Methods for Convection-Diffusion Equations by Volker John and Petr Knobloch.
template <int dim>
double compute_stabilization_delta(const double hk,
const double pk)
{
const double Peclet = dir.
norm() * hk / (2.0 *
eps * pk);
return hk / (2.0 * dir.
norm() * pk) * (
coth - 1.0 / Peclet);
}
numbers::NumberTraits< Number >::real_type norm() const
Expression coth(const Expression &x)
::VectorizedArray< Number, width > exp(const ::VectorizedArray< Number, width > &)
AdvectionProlem class
This is the main class of the program, and should look very similar to step-16. The major difference is that, since we are defining our multigrid smoother at runtime, we choose to define a function create_smoother() and a class object mg_smoother which is a std::unique_ptr to a smoother that is derived from MGSmoother. Note that for smoothers derived from RelaxationBlock, we must include a smoother_data object for each level. This will contain information about the cell ordering and the method of inverting cell matrices.
template <int dim>
class AdvectionProblem
{
public:
AdvectionProblem(
const Settings &settings);
private:
void setup_system();
template <class IteratorType>
void assemble_cell(const IteratorType &cell,
ScratchData<dim> & scratch_data,
CopyData & copy_data);
void assemble_system_and_multigrid();
void setup_smoother();
void solve();
void refine_grid();
void output_results(const unsigned int cycle) const;
std::unique_ptr<MGSmoother<Vector<double>>> mg_smoother;
using SmootherType =
using SmootherAdditionalDataType = SmootherType::AdditionalData;
};
template <int dim>
AdvectionProblem<dim>::AdvectionProblem(
const Settings &settings)
, fe(settings.fe_degree)
, mapping(settings.fe_degree)
, settings(settings)
{
if (dim >= 2)
if (dim >= 3)
}
static ::ExceptionBase & ExcNotImplemented()
void run(const Iterator &begin, const typename identity< Iterator >::type &end, Worker worker, Copier copier, const ScratchData &sample_scratch_data, const CopyData &sample_copy_data, const unsigned int queue_length, const unsigned int chunk_size)
static constexpr double PI
::VectorizedArray< Number, width > cos(const ::VectorizedArray< Number, width > &)
::VectorizedArray< Number, width > sin(const ::VectorizedArray< Number, width > &)
const ::parallel::distributed::Triangulation< dim, spacedim > * triangulation
AdvectionProblem::setup_system()
Here we first set up the DoFHandler, AffineConstraints, and SparsityPattern objects for both active and multigrid level meshes.
We could renumber the active DoFs with the DoFRenumbering class, but the smoothers only act on multigrid levels and as such, this would not matter for the computations. Instead, we will renumber the DoFs on each multigrid level below.
template <int dim>
void AdvectionProblem<dim>::setup_system()
{
solution.reinit(dof_handler.
n_dofs());
system_rhs.reinit(dof_handler.
n_dofs());
constraints.clear();
mapping, dof_handler, 0, BoundaryValues<dim>(), constraints);
mapping, dof_handler, 1, BoundaryValues<dim>(), constraints);
constraints.close();
dsp,
constraints,
false);
sparsity_pattern.copy_from(dsp);
system_matrix.reinit(sparsity_pattern);
void distribute_dofs(const FiniteElement< dim, spacedim > &fe)
void distribute_mg_dofs()
types::global_dof_index n_dofs() const
void make_hanging_node_constraints(const DoFHandler< dim, spacedim > &dof_handler, AffineConstraints< number > &constraints)
void make_sparsity_pattern(const DoFHandler< dim, spacedim > &dof_handler, SparsityPatternType &sparsity_pattern, const AffineConstraints< number > &constraints=AffineConstraints< number >(), const bool keep_constrained_dofs=true, const types::subdomain_id subdomain_id=numbers::invalid_subdomain_id)
Having enumerated the global degrees of freedom as well as (in the last line above) the level degrees of freedom, let us renumber the level degrees of freedom to get a better smoother as explained in the introduction. The first block below renumbers DoFs on each level in downstream or upstream direction if needed. This is only necessary for point smoothers (SOR and Jacobi) as the block smoothers operate on cells (see create_smoother()). The blocks below then also implement random numbering.
if (settings.smoother_type == "SOR" || settings.smoother_type == "Jacobi")
{
if (settings.dof_renumbering ==
settings.dof_renumbering ==
Settings::DoFRenumberingStrategy::upstream)
{
(settings.dof_renumbering ==
Settings::DoFRenumberingStrategy::upstream ?
-1.0 :
1.0) *
advection_direction;
direction,
true);
}
else if (settings.dof_renumbering ==
{
}
else
}
The rest of the function just sets up data structures. The last lines of the code below is unlike the other GMG tutorials, as it sets up both the interface in and out matrices. We need this since our problem is non-symmetric.
mg_constrained_dofs.clear();
mg_constrained_dofs.initialize(dof_handler);
mg_constrained_dofs.make_zero_boundary_constraints(dof_handler, {0, 1});
mg_matrices.resize(0, n_levels - 1);
mg_matrices.clear_elements();
mg_interface_in.resize(0, n_levels - 1);
mg_interface_in.clear_elements();
mg_interface_out.resize(0, n_levels - 1);
mg_interface_out.clear_elements();
mg_sparsity_patterns.resize(0, n_levels - 1);
mg_interface_sparsity_patterns.resize(0, n_levels - 1);
{
{
mg_sparsity_patterns[
level].copy_from(dsp);
mg_matrices[
level].reinit(mg_sparsity_patterns[
level]);
}
{
mg_constrained_dofs,
dsp,
mg_interface_sparsity_patterns[
level].copy_from(dsp);
mg_interface_in[
level].reinit(mg_interface_sparsity_patterns[
level]);
mg_interface_out[
level].reinit(mg_interface_sparsity_patterns[
level]);
}
}
}
AdvectionProblem::assemble_cell()
Here we define the assembly of the linear system on each cell to be used by the mesh_loop() function below. This one function assembles the cell matrix for either an active or a level cell (whatever it is passed as its first argument), and only assembles a right-hand side if called with an active cell.
template <int dim>
template <class IteratorType>
void AdvectionProblem<dim>::assemble_cell(const IteratorType &cell,
ScratchData<dim> & scratch_data,
CopyData & copy_data)
{
copy_data.level = cell->level();
const unsigned int dofs_per_cell =
scratch_data.fe_values.get_fe().n_dofs_per_cell();
copy_data.dofs_per_cell = dofs_per_cell;
copy_data.cell_matrix.reinit(dofs_per_cell, dofs_per_cell);
const unsigned int n_q_points =
scratch_data.fe_values.get_quadrature().size();
if (cell->is_level_cell() == false)
copy_data.cell_rhs.reinit(dofs_per_cell);
copy_data.local_dof_indices.resize(dofs_per_cell);
cell->get_active_or_mg_dof_indices(copy_data.local_dof_indices);
scratch_data.fe_values.reinit(cell);
RightHandSide<dim> right_hand_side;
std::vector<double> rhs_values(n_q_points);
right_hand_side.value_list(scratch_data.fe_values.get_quadrature_points(),
rhs_values);
If we are using streamline diffusion we must add its contribution to both the cell matrix and the cell right-hand side. If we are not using streamline diffusion, setting \delta=0 negates this contribution below and we are left with the standard, Galerkin finite element assembly.
const double delta = (settings.with_streamline_diffusion ?
compute_stabilization_delta(cell->diameter(),
settings.epsilon,
advection_direction,
settings.fe_degree) :
0.0);
for (unsigned int q_point = 0; q_point < n_q_points; ++q_point)
for (unsigned int i = 0; i < dofs_per_cell; ++i)
{
for (unsigned int j = 0; j < dofs_per_cell; ++j)
{
The assembly of the local matrix has two parts. First the Galerkin contribution:
copy_data.cell_matrix(i, j) +=
(settings.epsilon *
scratch_data.fe_values.shape_grad(i, q_point) *
scratch_data.fe_values.shape_grad(j, q_point) *
scratch_data.fe_values.JxW(q_point)) +
(scratch_data.fe_values.shape_value(i, q_point) *
(advection_direction *
scratch_data.fe_values.shape_grad(j, q_point)) *
scratch_data.fe_values.JxW(q_point))
and then the streamline diffusion contribution:
+ delta *
(advection_direction *
scratch_data.fe_values.shape_grad(j, q_point)) *
(advection_direction *
scratch_data.fe_values.shape_grad(i, q_point)) *
scratch_data.fe_values.JxW(q_point) -
delta * settings.epsilon *
trace(scratch_data.fe_values.shape_hessian(j, q_point)) *
(advection_direction *
scratch_data.fe_values.shape_grad(i, q_point)) *
scratch_data.fe_values.JxW(q_point);
}
if (cell->is_level_cell() == false)
{
constexpr Number trace(const SymmetricTensor< 2, dim, Number > &d)
The same applies to the right hand side. First the Galerkin contribution:
copy_data.cell_rhs(i) +=
scratch_data.fe_values.shape_value(i, q_point) *
rhs_values[q_point] * scratch_data.fe_values.JxW(q_point)
and then the streamline diffusion contribution:
+ delta * rhs_values[q_point] * advection_direction *
scratch_data.fe_values.shape_grad(i, q_point) *
scratch_data.fe_values.JxW(q_point);
}
}
}
AdvectionProblem::assemble_system_and_multigrid()
Here we employ MeshWorker::mesh_loop() to go over cells and assemble the system_matrix, system_rhs, and all mg_matrices for us.
template <int dim>
void AdvectionProblem<dim>::assemble_system_and_multigrid()
{
const auto cell_worker_active =
ScratchData<dim> & scratch_data,
CopyData & copy_data) {
this->assemble_cell(cell, scratch_data, copy_data);
};
const auto copier_active = [&](const CopyData ©_data) {
constraints.distribute_local_to_global(copy_data.cell_matrix,
copy_data.cell_rhs,
copy_data.local_dof_indices,
system_matrix,
system_rhs);
};
cell_worker_active,
copier_active,
ScratchData<dim>(fe, fe.degree + 1),
CopyData(),
cell_iterator end() const
active_cell_iterator begin_active(const unsigned int level=0) const
void mesh_loop(const CellIteratorType &begin, const CellIteratorType &end, const CellWorkerFunctionType &cell_worker, const CopierType &copier, const ScratchData &sample_scratch_data, const CopyData &sample_copy_data, const AssembleFlags flags=assemble_own_cells, const BoundaryWorkerFunctionType &boundary_worker=BoundaryWorkerFunctionType(), const FaceWorkerFunctionType &face_worker=FaceWorkerFunctionType(), const unsigned int queue_length=2 *MultithreadInfo::n_threads(), const unsigned int chunk_size=8)
Unlike the constraints for the active level, we choose to create constraint objects for each multigrid level local to this function since they are never needed elsewhere in the program.
std::vector<AffineConstraints<double>> boundary_constraints(
{
IndexSet locally_owned_level_dof_indices;
dof_handler,
level, locally_owned_level_dof_indices);
boundary_constraints[
level].reinit(locally_owned_level_dof_indices);
boundary_constraints[
level].add_lines(
mg_constrained_dofs.get_refinement_edge_indices(
level));
boundary_constraints[
level].add_lines(
mg_constrained_dofs.get_boundary_indices(
level));
boundary_constraints[
level].close();
}
const auto cell_worker_mg =
[&](
const decltype(dof_handler.
begin_mg()) &cell,
ScratchData<dim> & scratch_data,
CopyData & copy_data) {
this->assemble_cell(cell, scratch_data, copy_data);
};
const auto copier_mg = [&](const CopyData ©_data) {
boundary_constraints[copy_data.level].distribute_local_to_global(
copy_data.cell_matrix,
copy_data.local_dof_indices,
mg_matrices[copy_data.level]);
level_cell_iterator begin_mg(const unsigned int level=0) const
If (i,j) is an interface_out dof pair, then (j,i) is an interface_in dof pair. Note: For interface_in, we load the transpose of the interface entries, i.e., the entry for dof pair (j,i) is stored in interface_in(i,j). This is an optimization for the symmetric case which allows only one matrix to be used when setting the edge_matrices in solve(). Here, however, since our problem is non-symmetric, we must store both interface_in and interface_out matrices.
for (unsigned int i = 0; i < copy_data.dofs_per_cell; ++i)
for (unsigned int j = 0; j < copy_data.dofs_per_cell; ++j)
if (mg_constrained_dofs.is_interface_matrix_entry(
copy_data.level,
copy_data.local_dof_indices[i],
copy_data.local_dof_indices[j]))
{
mg_interface_out[copy_data.level].add(
copy_data.local_dof_indices[i],
copy_data.local_dof_indices[j],
copy_data.cell_matrix(i, j));
mg_interface_in[copy_data.level].add(
copy_data.local_dof_indices[i],
copy_data.local_dof_indices[j],
copy_data.cell_matrix(j, i));
}
};
cell_worker_mg,
copier_mg,
ScratchData<dim>(fe, fe.degree + 1),
CopyData(),
}
level_cell_iterator end_mg(const unsigned int level) const
AdvectionProblem::setup_smoother()
Next, we set up the smoother based on the settings in the .prm file. The two options that are of significance is the number of pre- and post-smoothing steps on each level of the multigrid v-cycle and the relaxation parameter.
Since multiplicative methods tend to be more powerful than additive method, fewer smoothing steps are required to see convergence independent of mesh size. The same holds for block smoothers over point smoothers. This is reflected in the choice for the number of smoothing steps for each type of smoother below.
The relaxation parameter for point smoothers is chosen based on trial and error, and reflects values necessary to keep the iteration counts in the GMRES solve constant (or as close as possible) as we refine the mesh. The two values given for both "Jacobi" and "SOR" in the .prm files are for degree 1 and degree 3 finite elements. If the user wants to change to another degree, they may need to adjust these numbers. For block smoothers, this parameter has a more straightforward interpretation, namely that for additive methods in 2D, a DoF can have a repeated contribution from up to 4 cells, therefore we must relax these methods by 0.25 to compensate. This is not an issue for multiplicative methods as each cell's inverse application carries new information to all its DoFs.
Finally, as mentioned above, the point smoothers only operate on DoFs, and the block smoothers on cells, so only the block smoothers need to be given information regarding cell orderings. DoF ordering for point smoothers has already been taken care of in setup_system().
template <int dim>
void AdvectionProblem<dim>::setup_smoother()
{
if (settings.smoother_type == "SOR")
{
auto smoother =
std::make_unique<MGSmootherPrecondition<SparseMatrix<double>,
Smoother,
smoother->initialize(mg_matrices,
Smoother::AdditionalData(fe.degree == 1 ? 1.0 :
0.62));
smoother->set_steps(settings.smoothing_steps);
mg_smoother = std::move(smoother);
}
else if (settings.smoother_type == "Jacobi")
{
auto smoother =
std::make_unique<MGSmootherPrecondition<SparseMatrix<double>,
Smoother,
smoother->initialize(mg_matrices,
Smoother::AdditionalData(fe.degree == 1 ? 0.6667 :
0.47));
smoother->set_steps(settings.smoothing_steps);
mg_smoother = std::move(smoother);
}
else if (settings.smoother_type == "block SOR" ||
settings.smoother_type == "block Jacobi")
{
{
dof_handler,
smoother_data[
level].relaxation =
(settings.smoother_type == "block SOR" ? 1.0 : 0.25);
std::vector<unsigned int> ordered_indices;
switch (settings.dof_renumbering)
{
ordered_indices =
create_downstream_cell_ordering(dof_handler,
advection_direction,
break;
case Settings::DoFRenumberingStrategy::upstream:
ordered_indices =
create_downstream_cell_ordering(dof_handler,
-1.0 * advection_direction,
break;
ordered_indices =
create_random_cell_ordering(dof_handler,
level);
break;
break;
default:
break;
}
smoother_data[
level].order =
std::vector<std::vector<unsigned int>>(1, ordered_indices);
}
if (settings.smoother_type == "block SOR")
{
smoother->initialize(mg_matrices, smoother_data);
smoother->set_steps(settings.smoothing_steps);
mg_smoother = std::move(smoother);
}
else if (settings.smoother_type == "block Jacobi")
{
auto smoother = std::make_unique<
double,
smoother->initialize(mg_matrices, smoother_data);
smoother->set_steps(settings.smoothing_steps);
mg_smoother = std::move(smoother);
}
}
else
}
AdvectionProblem::solve()
Before we can solve the system, we must first set up the multigrid preconditioner. This requires the setup of the transfer between levels, the coarse matrix solver, and the smoother. This setup follows almost identically to step-16, the main difference being the various smoothers defined above and the fact that we need different interface edge matrices for in and out since our problem is non-symmetric. (In reality, for this tutorial these interface matrices are empty since we are only using global refinement, and thus have no refinement edges. However, we have still included both here since if one made the simple switch to an adaptively refined method, the program would still run correctly.)
The last thing to note is that since our problem is non-symmetric, we must use an appropriate Krylov subspace method. We choose here to use GMRES since it offers the guarantee of residual reduction in each iteration. The major disavantage of GMRES is that, for each iteration, the number of stored temporary vectors increases by one, and one also needs to compute a scalar product with all previously stored vectors. This is rather expensive. This requirement is relaxed by using the restarted GMRES method which puts a cap on the number of vectors we are required to store at any one time (here we restart after 50 temporary vectors, or 48 iterations). This then has the disadvantage that we lose information we have gathered throughout the iteration and therefore we could see slower convergence. As a consequence, where to restart is a question of balancing memory consumption, CPU effort, and convergence speed. However, the goal of this tutorial is to have very low iteration counts by using a powerful GMG preconditioner, so we have picked the restart length such that all of the results shown below converge prior to restart happening, and thus we have a standard GMRES method. If the user is interested, another suitable method offered in deal.II would be BiCGStab.
template <int dim>
void AdvectionProblem<dim>::solve()
{
const unsigned int max_iters = 200;
const double solve_tolerance = 1
e-8 * system_rhs.l2_norm();
SolverControl solver_control(max_iters, solve_tolerance,
true,
true);
solver_control.enable_history_data();
Transfer mg_transfer(mg_constrained_dofs);
mg_transfer.build(dof_handler);
setup_smoother();
mg_matrix.initialize(mg_matrices);
mg_interface_matrix_in.initialize(mg_interface_in);
mg_interface_matrix_out.initialize(mg_interface_out);
mg_matrix, coarse_grid_solver, mg_transfer, *mg_smoother, *mg_smoother);
mg.set_edge_matrices(mg_interface_matrix_out, mg_interface_matrix_in);
mg_transfer);
std::cout << " Solving with GMRES to tol " << solve_tolerance << "..."
<< std::endl;
solver.solve(system_matrix, solution, system_rhs, preconditioner);
std::cout << " converged in " << solver_control.last_step()
<< " iterations"
constraints.distribute(solution);
mg_smoother.release();
}
void copy_from(const MatrixType &)
void initialize(const FullMatrix< number > &A)
double last_wall_time() const
AdvectionProblem::output_results()
The final function of interest generates graphical output. Here we output the solution and cell ordering in a .vtu format.
At the top of the function, we generate an index for each cell to visualize the ordering used by the smoothers. Note that we do this only for the active cells instead of the levels, where the smoothers are actually used. For the point smoothers we renumber DoFs instead of cells, so this is only an approximation of what happens in reality. Finally, the random ordering is not the random ordering we actually use (see create_smoother() for that).
The (integer) ordering of cells is then copied into a (floating point) vector for graphical output.
template <int dim>
void AdvectionProblem<dim>::output_results(const unsigned int cycle) const
{
{
std::vector<unsigned int> ordered_indices;
switch (settings.dof_renumbering)
{
ordered_indices =
create_downstream_cell_ordering(dof_handler, advection_direction);
break;
case Settings::DoFRenumberingStrategy::upstream:
ordered_indices =
create_downstream_cell_ordering(dof_handler,
-1.0 * advection_direction);
break;
ordered_indices = create_random_cell_ordering(dof_handler);
break;
ordered_indices[i] = i;
break;
default:
break;
}
cell_indices(ordered_indices[i]) = static_cast<double>(i);
}
unsigned int n_active_cells(const internal::TriangulationImplementation::NumberCache< 1 > &c)
The remainder of the function is then straightforward, given previous tutorial programs:
const std::string filename =
std::ofstream output(filename.c_str());
}
void attach_dof_handler(const DoFHandlerType &)
void add_data_vector(const VectorType &data, const std::vector< std::string > &names, const DataVectorType type=type_automatic, const std::vector< DataComponentInterpretation::DataComponentInterpretation > &data_component_interpretation=std::vector< DataComponentInterpretation::DataComponentInterpretation >())
virtual void build_patches(const unsigned int n_subdivisions=0)
void write_vtu(std::ostream &out) const
std::string int_to_string(const unsigned int value, const unsigned int digits=numbers::invalid_unsigned_int)
As in most tutorials, this function creates/refines the mesh and calls the various functions defined above to set up, assemble, solve, and output the results.
In cycle zero, we generate the mesh for the on the square [-1,1]^dim with a hole of radius 3/10 units centered at the origin. For objects with manifold_id equal to one (namely, the faces adjacent to the hole), we assign a spherical manifold.
template <int dim>
{
for (unsigned int cycle = 0; cycle < (settings.fe_degree == 1 ? 7 : 5);
++cycle)
{
std::cout << " Cycle " << cycle << ':' << std::endl;
if (cycle == 0)
{
0.3,
1.0);
}
setup_system();
std::cout << " Number of active cells: "
std::cout << " Number of degrees of freedom: "
<< dof_handler.
n_dofs() << std::endl;
assemble_system_and_multigrid();
solve();
if (settings.output)
output_results(cycle);
std::cout << std::endl;
}
}
}
void hyper_cube_with_cylindrical_hole(Triangulation< dim > &triangulation, const double inner_radius=.25, const double outer_radius=.5, const double L=.5, const unsigned int repetitions=1, const bool colorize=false)
The main function
Finally, the main function is like most tutorials. The only interesting bit is that we require the user to pass a .prm file as a sole command line argument. If no parameter file is given, the program will output the contents of a sample parameter file with all default values to the screen that the user can then copy and paste into their own .prm file.
int main(int argc, char *argv[])
{
try
{
settings.get_parameters((argc > 1) ? (argv[1]) : "");
Step63::AdvectionProblem<2> advection_problem_2d(settings);
advection_problem_2d.run();
}
catch (std::exception &exc)
{
std::cerr << std::endl
<< std::endl
<< "----------------------------------------------------"
<< std::endl;
std::cerr << "Exception on processing: " << std::endl
<< exc.what() << std::endl
<< "Aborting!" << std::endl
<< "----------------------------------------------------"
<< std::endl;
return 1;
}
catch (...)
{
std::cerr << std::endl
<< std::endl
<< "----------------------------------------------------"
<< std::endl;
std::cerr << "Unknown exception!" << std::endl
<< "Aborting!" << std::endl
<< "----------------------------------------------------"
<< std::endl;
return 1;
}
return 0;
}
Results
GMRES Iteration Numbers
The major advantage for GMG is that it is an \mathcal{O}(n) method, that is, the complexity of the problem increases linearly with the problem size. To show then that the linear solver presented in this tutorial is in fact \mathcal{O}(n), all one needs to do is show that the iteration counts for the GMRES solve stay roughly constant as we refine the mesh.
Each of the following tables gives the GMRES iteration counts to reduce the initial residual by a factor of 10^8. We selected a sufficient number of smoothing steps (based on the method) to get iteration numbers independent of mesh size. As can be seen from the tables below, the method is indeed \mathcal{O}(n).
DoF/Cell Renumbering
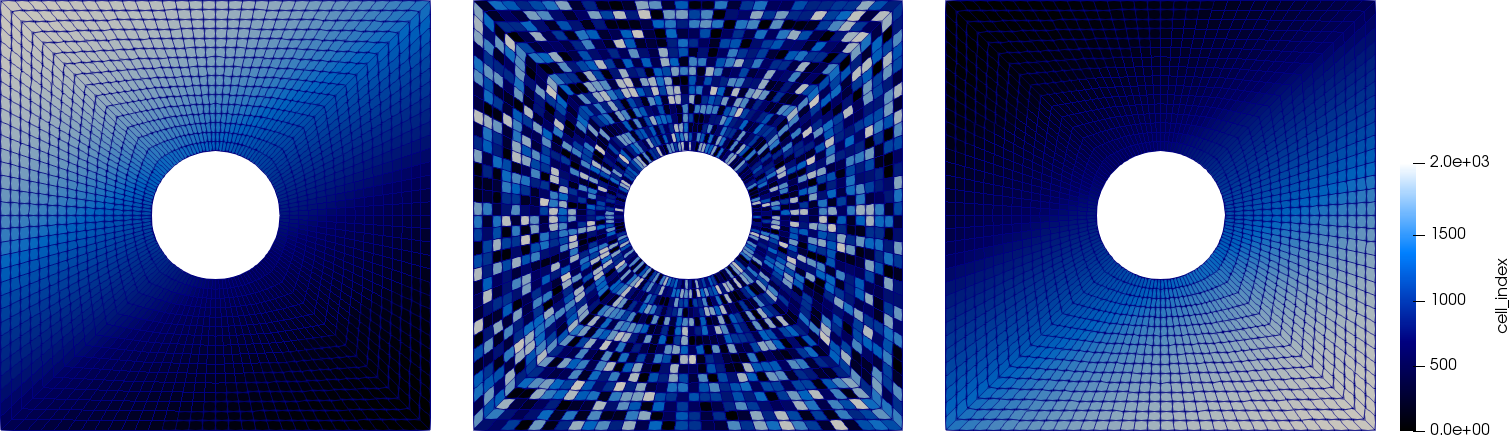
The point-wise smoothers ("Jacobi" and "SOR") get applied in the order the DoFs are numbered on each level. We can influence this using the DoFRenumbering namespace. The block smoothers are applied based on the ordering we set in setup_smoother(). We can visualize this numbering. The following pictures show the cell numbering of the active cells in downstream, random, and upstream numbering (left to right):
Let us start with the additive smoothers. The following table shows the number of iterations necessary to obtain convergence from GMRES:
| | Q_1 | Smoother (smoothing steps) |
| | | Jacobi (6) | | Block Jacobi (3) |
| | | Renumbering Strategy | | Renumbering Strategy |
| Cells | | DoFs | Downstream | Random | Upstream | | Downstream | Random | Upstream |
| 32 | | 48 | 3 | 3 | 3 | | 3 | 3 | 3 |
| 128 | | 160 | 6 | 6 | 6 | | 6 | 6 | 6 |
| 512 | | 576 | 11 | 11 | 11 | | 9 | 9 | 9 |
| 2048 | | 2176 | 15 | 15 | 15 | | 13 | 13 | 13 |
| 8192 | | 8448 | 18 | 18 | 18 | | 15 | 15 | 15 |
| 32768 | | 33280 | 20 | 20 | 20 | | 16 | 16 | 16 |
| 131072 | | 132096 | 20 | 20 | 20 | | 16 | 16 | 16 |
We see that renumbering the DoFs/cells has no effect on convergence speed. This is because these smoothers compute operations on each DoF (point-smoother) or cell (block-smoother) independently and add up the results. Since we can define these smoothers as an application of a sum of matrices, and matrix addition is commutative, the order at which we sum the different components will not affect the end result.
On the other hand, the situation is different for multiplicative smoothers:
| | Q_1 | Smoother (smoothing steps) |
| | | SOR (3) | | Block SOR (1) |
| | | Renumbering Strategy | | Renumbering Strategy |
| Cells | | DoFs | Downstream | Random | Upstream | | Downstream | Random | Upstream |
| 32 | | 48 | 2 | 2 | 3 | | 2 | 2 | 3 |
| 128 | | 160 | 5 | 5 | 7 | | 5 | 5 | 7 |
| 512 | | 576 | 7 | 9 | 11 | | 7 | 7 | 12 |
| 2048 | | 2176 | 10 | 12 | 15 | | 8 | 10 | 17 |
| 8192 | | 8448 | 11 | 15 | 19 | | 10 | 11 | 20 |
| 32768 | | 33280 | 12 | 16 | 20 | | 10 | 12 | 21 |
| 131072 | | 132096 | 12 | 16 | 19 | | 11 | 12 | 21 |
Here, we can speed up convergence by renumbering the DoFs/cells in the advection direction, and similarly, we can slow down convergence if we do the renumbering in the opposite direction. This is because advection-dominated problems have a directional flow of information (in the advection direction) which, given the right renumbering of DoFs/cells, multiplicative methods are able to capture.
This feature of multiplicative methods is, however, dependent on the value of \varepsilon. As we increase \varepsilon and the problem becomes more diffusion-dominated, we have a more uniform propagation of information over the mesh and there is a diminished advantage for renumbering in the advection direction. On the opposite end, in the extreme case of \varepsilon=0 (advection-only), we have a 1st-order PDE and multiplicative methods with the right renumbering become effective solvers: A correct downstream numbering may lead to methods that require only a single iteration because information can be propagated from the inflow boundary downstream, with no information transport in the opposite direction. (Note, however, that in the case of \varepsilon=0, special care must be taken for the boundary conditions in this case).
Point vs. block smoothers
We will limit the results to runs using the downstream renumbering. Here is a cross comparison of all four smoothers for both Q_1 and Q_3 elements:
| | Q_1 | Smoother (smoothing steps) | | Q_3 | Smoother (smoothing steps) |
| Cells | | DoFs | Jacobi (6) | Block Jacobi (3) | SOR (3) | Block SOR (1) | | DoFs | Jacobi (6) | Block Jacobi (3) | SOR (3) | Block SOR (1) |
| 32 | | 48 | 3 | 3 | 2 | 2 | | 336 | 15 | 14 | 15 | 6 |
| 128 | | 160 | 6 | 6 | 5 | 5 | | 1248 | 23 | 18 | 21 | 9 |
| 512 | | 576 | 11 | 9 | 7 | 7 | | 4800 | 29 | 21 | 28 | 9 |
| 2048 | | 2176 | 15 | 13 | 10 | 8 | | 18816 | 33 | 22 | 32 | 9 |
| 8192 | | 8448 | 18 | 15 | 11 | 10 | | 74496 | 35 | 22 | 34 | 10 |
| 32768 | | 33280 | 20 | 16 | 12 | 10 | |
| 131072 | | 132096 | 20 | 16 | 12 | 11 | |
We see that for Q_1, both multiplicative smoothers require a smaller combination of smoothing steps and iteration counts than either additive smoother. However, when we increase the degree to a Q_3 element, there is a clear advantage for the block smoothers in terms of the number of smoothing steps and iterations required to solve. Specifically, the block SOR smoother gives constant iteration counts over the degree, and the block Jacobi smoother only sees about a 38% increase in iterations compared to 75% and 183% for Jacobi and SOR respectively.
Cost
Iteration counts do not tell the full story in the optimality of a one smoother over another. Obviously we must examine the cost of an iteration. Block smoothers here are at a disadvantage as they are having to construct and invert a cell matrix for each cell. Here is a comparison of solve times for a Q_3 element with 74,496 DoFs:
| Q_3 | Smoother (smoothing steps) |
| DoFs | Jacobi (6) | Block Jacobi (3) | SOR (3) | Block SOR (1) |
| 74496 | 0.68s | 5.82s | 1.18s | 1.02s |
The smoother that requires the most iterations (Jacobi) actually takes the shortest time (roughly 2/3 the time of the next fastest method). This is because all that is required to apply a Jacobi smoothing step is multiplication by a diagonal matrix which is very cheap. On the other hand, while SOR requires over 3x more iterations (each with 3x more smoothing steps) than block SOR, the times are roughly equivalent, implying that a smoothing step of block SOR is roughly 9x slower than a smoothing step of SOR. Lastly, block Jacobi is almost 6x more expensive than block SOR, which intuitively makes sense from the fact that 1 step of each method has the same cost (inverting the cell matrices and either adding or multiply them together), and block Jacobi has 3 times the number of smoothing steps per iteration with 2 times the iterations.
Additional points
There are a few more important points to mention:
-
For a mesh distributed in parallel, multiplicative methods cannot be executed over the entire domain. This is because they operate one cell at a time, and downstream cells can only be handled once upstream cells have already been done. This is fine on a single processor: The processor just goes through the list of cells one after the other. However, in parallel, it would imply that some processors are idle because upstream processors have not finished doing the work on cells upstream from the ones owned by the current processor. Once the upstream processors are done, the downstream ones can start, but by that time the upstream processors have no work left. In other words, most of the time during these smoother steps, most processors are in fact idle. This is not how one obtains good parallel scalability!
One can use a hybrid method where a multiplicative smoother is applied on each subdomain, but as you increase the number of subdomains, the method approaches the behavior of an additive method. This is a major disadvantage to these methods.
-
Current research into block smoothers suggest that soon we will be able to compute the inverse of the cell matrices much cheaper than what is currently being done inside deal.II. This research is based on the fast diagonalization method (dating back to the 1960s) and has been used in the spectral community for around 20 years (see, e.g., Hybrid Multigrid/Schwarz Algorithms for the Spectral Element Method by Lottes and Fischer). There are currently efforts to generalize these methods to DG and make them more robust. Also, it seems that one should be able to take advantage of matrix-free implementations and the fact that, in the interior of the domain, cell matrices tend to look very similar, allowing fewer matrix inverse computations.
Combining 1. and 2. gives a good reason for expecting that a method like block Jacobi could become very powerful in the future, even though currently for these examples it is quite slow.
Possibilities for extensions
Constant iterations for Q5
Change the number of smoothing steps and the smoother relaxation parameter (set in Smoother::AdditionalData() inside create_smoother(), only necessary for point smoothers) so that we maintain a constant number of iterations for a Q_5 element.
Effectiveness of renumbering for changing epsilon
Increase/decrease the parameter "Epsilon" in the .prm files of the multiplicative methods and observe for which values renumbering no longer influences convergence speed.
Mesh adaptivity
The code is set up to work correctly with an adaptively refined mesh (the interface matrices are created and set). Devise a suitable refinement criterium or try the KellyErrorEstimator class.
The plain program
#include <algorithm>
#include <fstream>
#include <iostream>
#include <random>
namespace Step63
{
template <int dim>
struct ScratchData
{
const unsigned int quadrature_degree)
: fe_values(fe,
QGauss<dim>(quadrature_degree),
{}
ScratchData(const ScratchData<dim> &scratch_data)
: fe_values(scratch_data.fe_values.get_fe(),
scratch_data.fe_values.get_quadrature(),
{}
};
struct CopyData
{
CopyData() = default;
unsigned int dofs_per_cell;
std::vector<types::global_dof_index> local_dof_indices;
};
{
enum DoFRenumberingStrategy
{
upstream,
};
void get_parameters(const std::string &prm_filename);
unsigned int fe_degree;
std::string smoother_type;
unsigned int smoothing_steps;
DoFRenumberingStrategy dof_renumbering;
bool with_streamline_diffusion;
bool output;
};
void Settings::get_parameters(const std::string &prm_filename)
{
"0.005",
"Diffusion parameter");
"1",
"Finite Element degree");
"block SOR",
"Select smoother: SOR|Jacobi|block SOR|block Jacobi");
"2",
"Number of smoothing steps");
"DoF renumbering",
"downstream",
"Select DoF renumbering: none|downstream|upstream|random");
"true",
"Enable streamline diffusion stabilization: true|false");
"true",
"Generate graphical output: true|false");
if (prm_filename.empty())
{
false,
ExcMessage(
"Please pass a .prm file as the first argument!"));
}
smoother_type = prm.
get(
"Smoother type");
const std::string renumbering = prm.
get(
"DoF renumbering");
if (renumbering == "none")
else if (renumbering == "downstream")
else if (renumbering == "upstream")
dof_renumbering = DoFRenumberingStrategy::upstream;
else if (renumbering == "random")
else
"an invalid value."));
with_streamline_diffusion = prm.
get_bool(
"With streamline diffusion");
}
template <int dim>
std::vector<unsigned int>
const unsigned int level)
{
std::vector<typename DoFHandler<dim>::level_cell_iterator> ordered_cells;
ordered_cells.push_back(cell);
const DoFRenumbering::
CompareDownstream<typename DoFHandler<dim>::level_cell_iterator, dim>
comparator(direction);
std::sort(ordered_cells.begin(), ordered_cells.end(), comparator);
std::vector<unsigned> ordered_indices;
for (const auto &cell : ordered_cells)
ordered_indices.push_back(cell->index());
return ordered_indices;
}
template <int dim>
std::vector<unsigned int>
{
std::vector<typename DoFHandler<dim>::active_cell_iterator> ordered_cells;
ordered_cells.push_back(cell);
const DoFRenumbering::
CompareDownstream<typename DoFHandler<dim>::active_cell_iterator, dim>
comparator(direction);
std::sort(ordered_cells.begin(), ordered_cells.end(), comparator);
std::vector<unsigned int> ordered_indices;
for (const auto &cell : ordered_cells)
ordered_indices.push_back(cell->index());
return ordered_indices;
}
template <int dim>
std::vector<unsigned int>
const unsigned int level)
{
std::vector<unsigned int> ordered_cells;
ordered_cells.push_back(cell->index());
std::mt19937 random_number_generator;
std::shuffle(ordered_cells.begin(),
ordered_cells.end(),
random_number_generator);
return ordered_cells;
}
template <int dim>
std::vector<unsigned int>
{
std::vector<unsigned int> ordered_cells;
ordered_cells.push_back(cell->index());
std::mt19937 random_number_generator;
std::shuffle(ordered_cells.begin(),
ordered_cells.end(),
random_number_generator);
return ordered_cells;
}
template <int dim>
class RightHandSide :
public Function<dim>
{
public:
const unsigned int component = 0) const override;
virtual void value_list(
const std::vector<
Point<dim>> &points,
const unsigned int component = 0) const override;
};
template <int dim>
double RightHandSide<dim>::value(
const Point<dim> &,
const unsigned int component) const
{
(void)component;
return 0.0;
}
template <int dim>
void RightHandSide<dim>::value_list(
const std::vector<
Point<dim>> &points,
const unsigned int component) const
{
for (unsigned int i = 0; i < points.size(); ++i)
values[i] = RightHandSide<dim>::value(points[i], component);
}
template <int dim>
class BoundaryValues :
public Function<dim>
{
public:
const unsigned int component = 0) const override;
virtual void value_list(
const std::vector<
Point<dim>> &points,
const unsigned int component = 0) const override;
};
template <int dim>
double BoundaryValues<dim>::value(
const Point<dim> & p,
const unsigned int component) const
{
(void)component;
{
return 1.0;
}
else
{
return 0.0;
}
}
template <int dim>
void BoundaryValues<dim>::value_list(
const std::vector<
Point<dim>> &points,
const unsigned int component) const
{
for (unsigned int i = 0; i < points.size(); ++i)
values[i] = BoundaryValues<dim>::value(points[i], component);
}
template <int dim>
double compute_stabilization_delta(const double hk,
const double pk)
{
const double Peclet = dir.
norm() * hk / (2.0 *
eps * pk);
return hk / (2.0 * dir.
norm() * pk) * (
coth - 1.0 / Peclet);
}
template <int dim>
class AdvectionProblem
{
public:
AdvectionProblem(
const Settings &settings);
private:
void setup_system();
template <class IteratorType>
void assemble_cell(const IteratorType &cell,
ScratchData<dim> & scratch_data,
CopyData & copy_data);
void assemble_system_and_multigrid();
void setup_smoother();
void solve();
void refine_grid();
void output_results(const unsigned int cycle) const;
std::unique_ptr<MGSmoother<Vector<double>>> mg_smoother;
using SmootherType =
using SmootherAdditionalDataType = SmootherType::AdditionalData;
};
template <int dim>
AdvectionProblem<dim>::AdvectionProblem(
const Settings &settings)
, fe(settings.fe_degree)
, mapping(settings.fe_degree)
, settings(settings)
{
if (dim >= 2)
if (dim >= 3)
}
template <int dim>
void AdvectionProblem<dim>::setup_system()
{
solution.reinit(dof_handler.
n_dofs());
system_rhs.reinit(dof_handler.
n_dofs());
constraints.clear();
mapping, dof_handler, 0, BoundaryValues<dim>(), constraints);
mapping, dof_handler, 1, BoundaryValues<dim>(), constraints);
constraints.close();
dsp,
constraints,
false);
sparsity_pattern.copy_from(dsp);
system_matrix.reinit(sparsity_pattern);
if (settings.smoother_type == "SOR" || settings.smoother_type == "Jacobi")
{
if (settings.dof_renumbering ==
settings.dof_renumbering ==
Settings::DoFRenumberingStrategy::upstream)
{
(settings.dof_renumbering ==
Settings::DoFRenumberingStrategy::upstream ?
-1.0 :
1.0) *
advection_direction;
direction,
true);
}
else if (settings.dof_renumbering ==
{
}
else
}
mg_constrained_dofs.clear();
mg_constrained_dofs.initialize(dof_handler);
mg_constrained_dofs.make_zero_boundary_constraints(dof_handler, {0, 1});
mg_matrices.resize(0, n_levels - 1);
mg_matrices.clear_elements();
mg_interface_in.resize(0, n_levels - 1);
mg_interface_in.clear_elements();
mg_interface_out.resize(0, n_levels - 1);
mg_interface_out.clear_elements();
mg_sparsity_patterns.resize(0, n_levels - 1);
mg_interface_sparsity_patterns.resize(0, n_levels - 1);
{
{
mg_sparsity_patterns[
level].copy_from(dsp);
mg_matrices[
level].reinit(mg_sparsity_patterns[
level]);
}
{
mg_constrained_dofs,
dsp,
mg_interface_sparsity_patterns[
level].copy_from(dsp);
mg_interface_in[
level].reinit(mg_interface_sparsity_patterns[
level]);
mg_interface_out[
level].reinit(mg_interface_sparsity_patterns[
level]);
}
}
}
template <int dim>
template <class IteratorType>
void AdvectionProblem<dim>::assemble_cell(const IteratorType &cell,
ScratchData<dim> & scratch_data,
CopyData & copy_data)
{
copy_data.level = cell->level();
const unsigned int dofs_per_cell =
scratch_data.fe_values.get_fe().n_dofs_per_cell();
copy_data.dofs_per_cell = dofs_per_cell;
copy_data.cell_matrix.reinit(dofs_per_cell, dofs_per_cell);
const unsigned int n_q_points =
scratch_data.fe_values.get_quadrature().size();
if (cell->is_level_cell() == false)
copy_data.cell_rhs.reinit(dofs_per_cell);
copy_data.local_dof_indices.resize(dofs_per_cell);
cell->get_active_or_mg_dof_indices(copy_data.local_dof_indices);
scratch_data.fe_values.reinit(cell);
RightHandSide<dim> right_hand_side;
std::vector<double> rhs_values(n_q_points);
right_hand_side.value_list(scratch_data.fe_values.get_quadrature_points(),
rhs_values);
const double delta = (settings.with_streamline_diffusion ?
compute_stabilization_delta(cell->diameter(),
settings.epsilon,
advection_direction,
settings.fe_degree) :
0.0);
for (unsigned int q_point = 0; q_point < n_q_points; ++q_point)
for (unsigned int i = 0; i < dofs_per_cell; ++i)
{
for (unsigned int j = 0; j < dofs_per_cell; ++j)
{
copy_data.cell_matrix(i, j) +=
(settings.epsilon *
scratch_data.fe_values.shape_grad(i, q_point) *
scratch_data.fe_values.shape_grad(j, q_point) *
scratch_data.fe_values.JxW(q_point)) +
(scratch_data.fe_values.shape_value(i, q_point) *
(advection_direction *
scratch_data.fe_values.shape_grad(j, q_point)) *
scratch_data.fe_values.JxW(q_point))
+ delta *
(advection_direction *
scratch_data.fe_values.shape_grad(j, q_point)) *
(advection_direction *
scratch_data.fe_values.shape_grad(i, q_point)) *
scratch_data.fe_values.JxW(q_point) -
delta * settings.epsilon *
trace(scratch_data.fe_values.shape_hessian(j, q_point)) *
(advection_direction *
scratch_data.fe_values.shape_grad(i, q_point)) *
scratch_data.fe_values.JxW(q_point);
}
if (cell->is_level_cell() == false)
{
copy_data.cell_rhs(i) +=
scratch_data.fe_values.shape_value(i, q_point) *
rhs_values[q_point] * scratch_data.fe_values.JxW(q_point)
+ delta * rhs_values[q_point] * advection_direction *
scratch_data.fe_values.shape_grad(i, q_point) *
scratch_data.fe_values.JxW(q_point);
}
}
}
template <int dim>
void AdvectionProblem<dim>::assemble_system_and_multigrid()
{
const auto cell_worker_active =
ScratchData<dim> & scratch_data,
CopyData & copy_data) {
this->assemble_cell(cell, scratch_data, copy_data);
};
const auto copier_active = [&](const CopyData ©_data) {
constraints.distribute_local_to_global(copy_data.cell_matrix,
copy_data.cell_rhs,
copy_data.local_dof_indices,
system_matrix,
system_rhs);
};
cell_worker_active,
copier_active,
ScratchData<dim>(fe, fe.degree + 1),
CopyData(),
std::vector<AffineConstraints<double>> boundary_constraints(
{
IndexSet locally_owned_level_dof_indices;
dof_handler,
level, locally_owned_level_dof_indices);
boundary_constraints[
level].reinit(locally_owned_level_dof_indices);
boundary_constraints[
level].add_lines(
mg_constrained_dofs.get_refinement_edge_indices(
level));
boundary_constraints[
level].add_lines(
mg_constrained_dofs.get_boundary_indices(
level));
boundary_constraints[
level].close();
}
const auto cell_worker_mg =
[&](
const decltype(dof_handler.
begin_mg()) &cell,
ScratchData<dim> & scratch_data,
CopyData & copy_data) {
this->assemble_cell(cell, scratch_data, copy_data);
};
const auto copier_mg = [&](const CopyData ©_data) {
boundary_constraints[copy_data.level].distribute_local_to_global(
copy_data.cell_matrix,
copy_data.local_dof_indices,
mg_matrices[copy_data.level]);
for (unsigned int i = 0; i < copy_data.dofs_per_cell; ++i)
for (unsigned int j = 0; j < copy_data.dofs_per_cell; ++j)
if (mg_constrained_dofs.is_interface_matrix_entry(
copy_data.level,
copy_data.local_dof_indices[i],
copy_data.local_dof_indices[j]))
{
mg_interface_out[copy_data.level].add(
copy_data.local_dof_indices[i],
copy_data.local_dof_indices[j],
copy_data.cell_matrix(i, j));
mg_interface_in[copy_data.level].add(
copy_data.local_dof_indices[i],
copy_data.local_dof_indices[j],
copy_data.cell_matrix(j, i));
}
};
cell_worker_mg,
copier_mg,
ScratchData<dim>(fe, fe.degree + 1),
CopyData(),
}
template <int dim>
void AdvectionProblem<dim>::setup_smoother()
{
if (settings.smoother_type == "SOR")
{
auto smoother =
std::make_unique<MGSmootherPrecondition<SparseMatrix<double>,
Smoother,
smoother->initialize(mg_matrices,
Smoother::AdditionalData(fe.degree == 1 ? 1.0 :
0.62));
smoother->set_steps(settings.smoothing_steps);
mg_smoother = std::move(smoother);
}
else if (settings.smoother_type == "Jacobi")
{
auto smoother =
std::make_unique<MGSmootherPrecondition<SparseMatrix<double>,
Smoother,
smoother->initialize(mg_matrices,
Smoother::AdditionalData(fe.degree == 1 ? 0.6667 :
0.47));
smoother->set_steps(settings.smoothing_steps);
mg_smoother = std::move(smoother);
}
else if (settings.smoother_type == "block SOR" ||
settings.smoother_type == "block Jacobi")
{
{
dof_handler,
smoother_data[
level].relaxation =
(settings.smoother_type == "block SOR" ? 1.0 : 0.25);
std::vector<unsigned int> ordered_indices;
switch (settings.dof_renumbering)
{
ordered_indices =
create_downstream_cell_ordering(dof_handler,
advection_direction,
break;
case Settings::DoFRenumberingStrategy::upstream:
ordered_indices =
create_downstream_cell_ordering(dof_handler,
-1.0 * advection_direction,
break;
ordered_indices =
create_random_cell_ordering(dof_handler,
level);
break;
break;
default:
break;
}
smoother_data[
level].order =
std::vector<std::vector<unsigned int>>(1, ordered_indices);
}
if (settings.smoother_type == "block SOR")
{
smoother->initialize(mg_matrices, smoother_data);
smoother->set_steps(settings.smoothing_steps);
mg_smoother = std::move(smoother);
}
else if (settings.smoother_type == "block Jacobi")
{
auto smoother = std::make_unique<
double,
smoother->initialize(mg_matrices, smoother_data);
smoother->set_steps(settings.smoothing_steps);
mg_smoother = std::move(smoother);
}
}
else
}
template <int dim>
void AdvectionProblem<dim>::solve()
{
const unsigned int max_iters = 200;
const double solve_tolerance = 1
e-8 * system_rhs.l2_norm();
SolverControl solver_control(max_iters, solve_tolerance,
true,
true);
solver_control.enable_history_data();
Transfer mg_transfer(mg_constrained_dofs);
mg_transfer.build(dof_handler);
setup_smoother();
mg_matrix.initialize(mg_matrices);
mg_interface_matrix_in.initialize(mg_interface_in);
mg_interface_matrix_out.initialize(mg_interface_out);
mg_matrix, coarse_grid_solver, mg_transfer, *mg_smoother, *mg_smoother);
mg.set_edge_matrices(mg_interface_matrix_out, mg_interface_matrix_in);
mg_transfer);
std::cout << " Solving with GMRES to tol " << solve_tolerance << "..."
<< std::endl;
solver.solve(system_matrix, solution, system_rhs, preconditioner);
std::cout << " converged in " << solver_control.last_step()
<< " iterations"
constraints.distribute(solution);
mg_smoother.release();
}
template <int dim>
void AdvectionProblem<dim>::output_results(const unsigned int cycle) const
{
{
std::vector<unsigned int> ordered_indices;
switch (settings.dof_renumbering)
{
ordered_indices =
create_downstream_cell_ordering(dof_handler, advection_direction);
break;
case Settings::DoFRenumberingStrategy::upstream:
ordered_indices =
create_downstream_cell_ordering(dof_handler,
-1.0 * advection_direction);
break;
ordered_indices = create_random_cell_ordering(dof_handler);
break;
ordered_indices[i] = i;
break;
default:
break;
}
cell_indices(ordered_indices[i]) = static_cast<double>(i);
}
const std::string filename =
std::ofstream output(filename.c_str());
}
template <int dim>
{
for (unsigned int cycle = 0; cycle < (settings.fe_degree == 1 ? 7 : 5);
++cycle)
{
std::cout << " Cycle " << cycle << ':' << std::endl;
if (cycle == 0)
{
0.3,
1.0);
}
setup_system();
std::cout << " Number of active cells: "
std::cout << " Number of degrees of freedom: "
<< dof_handler.
n_dofs() << std::endl;
assemble_system_and_multigrid();
solve();
if (settings.output)
output_results(cycle);
std::cout << std::endl;
}
}
}
int main(int argc, char *argv[])
{
try
{
settings.get_parameters((argc > 1) ? (argv[1]) : "");
Step63::AdvectionProblem<2> advection_problem_2d(settings);
advection_problem_2d.run();
}
catch (std::exception &exc)
{
std::cerr << std::endl
<< std::endl
<< "----------------------------------------------------"
<< std::endl;
std::cerr << "Exception on processing: " << std::endl
<< exc.what() << std::endl
<< "Aborting!" << std::endl
<< "----------------------------------------------------"
<< std::endl;
return 1;
}
catch (...)
{
std::cerr << std::endl
<< std::endl
<< "----------------------------------------------------"
<< std::endl;
std::cerr << "Unknown exception!" << std::endl
<< "Aborting!" << std::endl
<< "----------------------------------------------------"
<< std::endl;
return 1;
}
return 0;
}